Description
Features:
- One single reagent is able to transfect large plasmid, mRNA, siRNA, and/or other type of nucleic acids, which is best for co-transfection of different type and/or size of nucleic acids.
- Broad Spectrum DNA/siRNA delivery – one transfection reagent and protocol for a variety of cells.
- Transfection efficiency is close to that of Avalanche®-Omni Transfection Reagent (EZT-OMNI-1), but with much lower prices. Its price is only 1/3 that of L2K, and 1/3.4 that of Lipofectamine® 3000 (L3K).
- Highest value:® It is able to do 3000 x 24-well transfections/1.5 ml as compared to 750-1000 x 24-well transfections/1.5 ml for L2K
- Very low Cellular Toxicity because of its bio-degradability after endocytosis. Maintain cell density, reduce experimental biases.
- Same simple protocol as that of EZT-OMNI-1: Does not require removal of serum or culture medium and does not require washing or changing of® medium after introducing the reagent/DNA complex.
- High levels of recombinant protein production
- Ideal for high-throughput work
- A must-have transfection reagent for everyday use on commonly used cells
Data
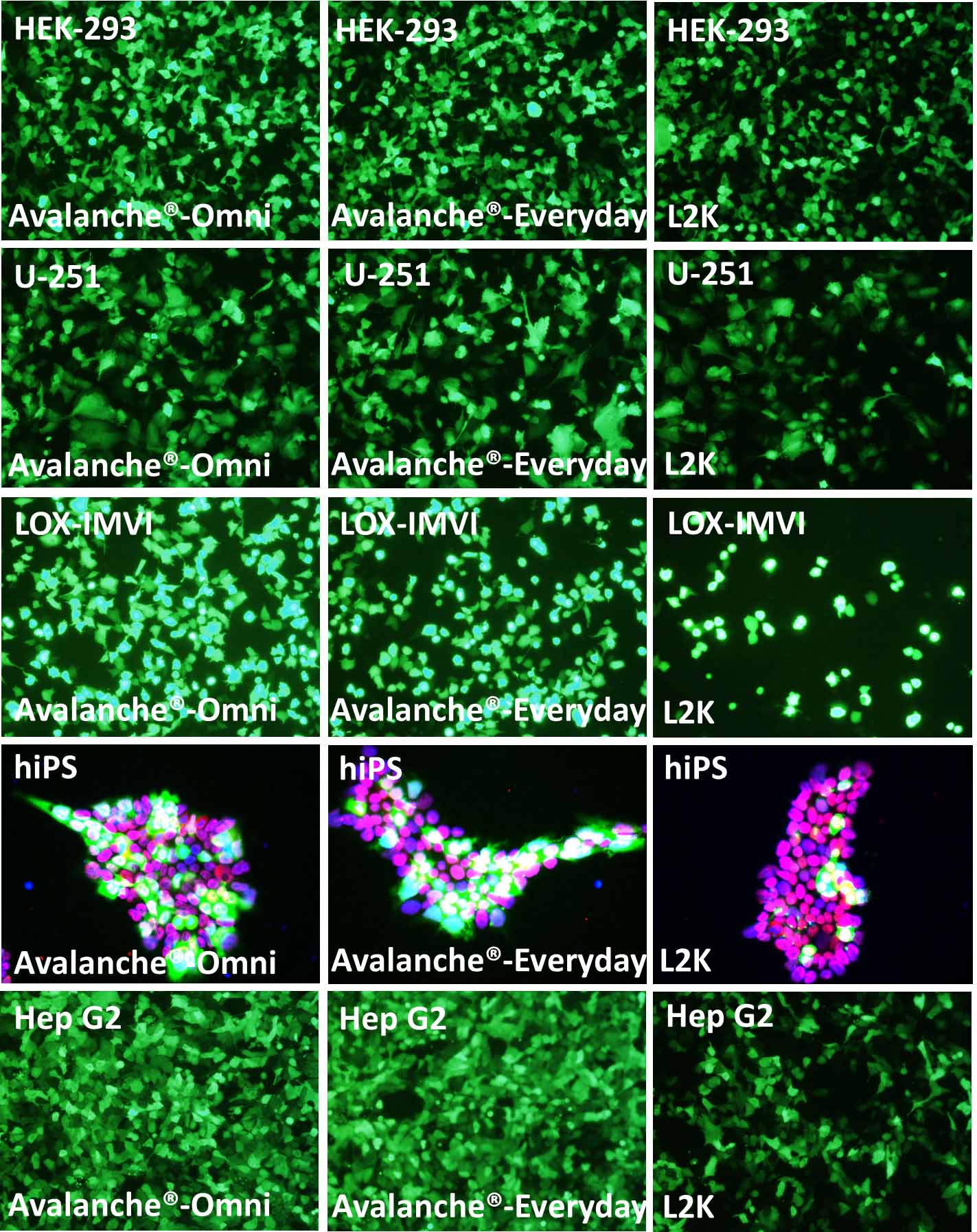 Figure 1. The performance of Avalanche®-Everyday Transfection Reagent (Avalanche®-Everyday) is very close to that of Avalanche®-Omni Transfection Reagent (Avalanche®-Omni). Cell line tested here are: HEK-293 (Human embryonic kidney cell line), U251 (Human glioblastoma cell), LOX-IMVI (Human melanoma cell), hiPS (Human induced pluripotent cell), and Hep G2 (Human hepatocellular carcinoma cell). The cells were transfected with enhanced green fluorescent protein (eGFP) expressing vector by using Avalanche®-Omni, Avalanche®-Everyday, and L2K. Regarding to hiPS cells, after transfection, hiPS cells were fixed with PFA, and stained with OCT-4 antibody (red) and DAPI (Blue). The pictures, which were the overlay of eGFP (green), Oct-4 (red), and DAPI (blue), showed that while all of the hiPS kept their pluripotency after transfection, Avalanche®-Omni and Avalanche®-Everyday showed great eGFP expression.
Figure 1. The performance of Avalanche®-Everyday Transfection Reagent (Avalanche®-Everyday) is very close to that of Avalanche®-Omni Transfection Reagent (Avalanche®-Omni). Cell line tested here are: HEK-293 (Human embryonic kidney cell line), U251 (Human glioblastoma cell), LOX-IMVI (Human melanoma cell), hiPS (Human induced pluripotent cell), and Hep G2 (Human hepatocellular carcinoma cell). The cells were transfected with enhanced green fluorescent protein (eGFP) expressing vector by using Avalanche®-Omni, Avalanche®-Everyday, and L2K. Regarding to hiPS cells, after transfection, hiPS cells were fixed with PFA, and stained with OCT-4 antibody (red) and DAPI (Blue). The pictures, which were the overlay of eGFP (green), Oct-4 (red), and DAPI (blue), showed that while all of the hiPS kept their pluripotency after transfection, Avalanche®-Omni and Avalanche®-Everyday showed great eGFP expression.
Avalanche®-Everyday and Avalanche®-Omni are able to achieved more than 90% knockdown of endogenous gene expression in a variety of cell lines and primary cells.
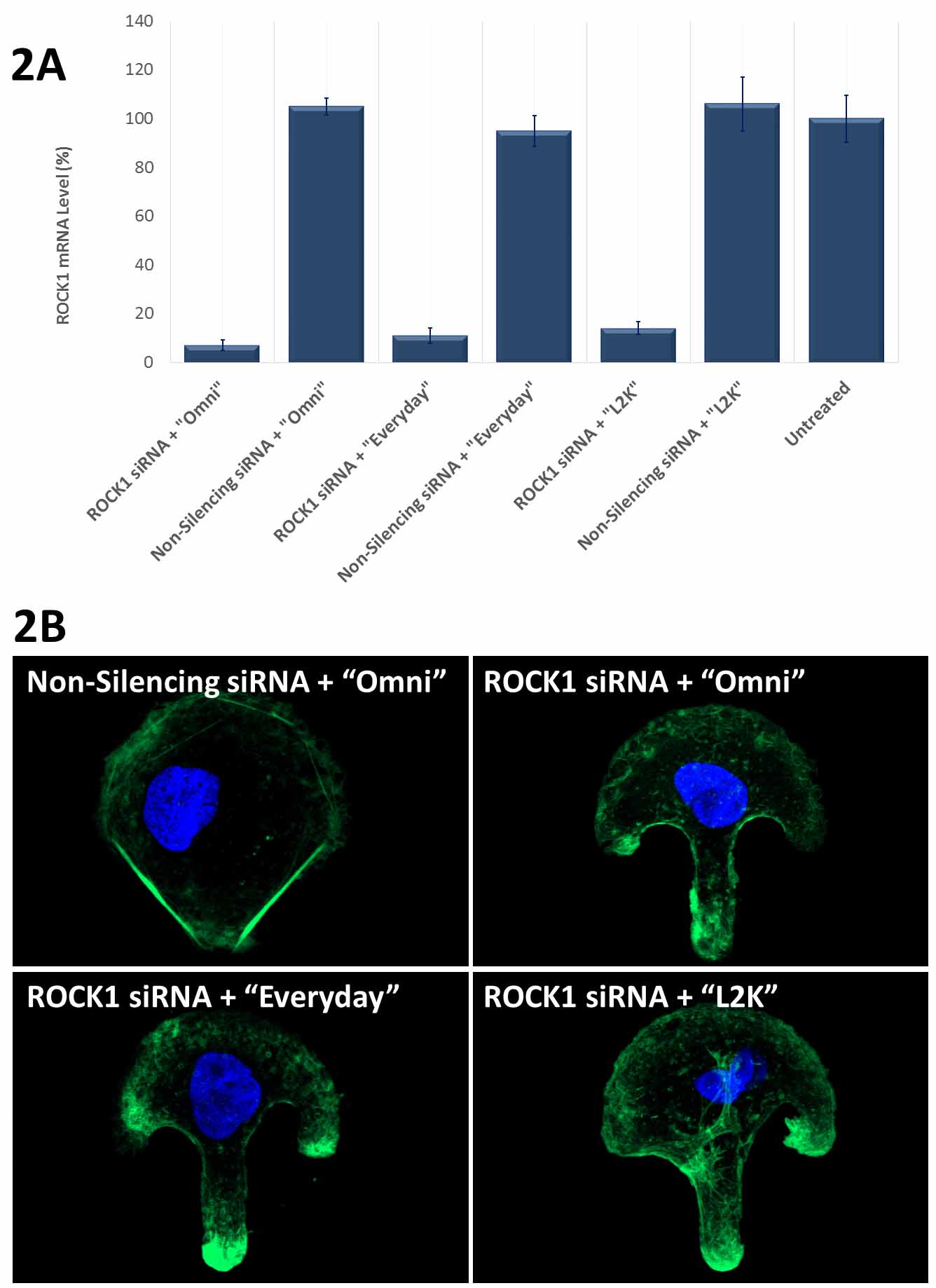 Figure 2. Avalanche®-Everyday Transfection Reagent (“Everyday”) and Avalanche®-Omni Transfection Reagent (“Omni”) achieved more than 90% knockdown of endogenous gene expression. (2A). ROCK1 mRNA levels were quantified using qPCR in Hela cells transfected with Target-specific siRNA duplexes (10nM) for human ROCK1 gene or non-silencing siRNA by using “Omni”, “Everyday”, and Lipofectamine® 2000 (“L2K”). Data were normalized against the 18S rRNA signal. Control Samples were either mock-transected or untreated. Values are normalized to untreated sample. Data are means®®SD (n=3). (B) Actin stress fibers were stained with FITC-labelled phalloidin on the cells in (A) cultured on fibronectin-coated micropattern. Confocal Microscope revealed that the cells transfected with ROCK1 siRNA showed disrupted stress fiber pattern as compared to the cells transfected with non-silencing siRNA using the same transfection reagent “Omni”. The use of “Everyday” showed similar disrupted stress fiber pattern as that of “Omni”, while the use of “L2K” only showed slightly disrupted stress fiber pattern. Lipofectamine® is a trademark of Life Technologies Corporation. Avalanche® is a trademark of EZ Biosystems® LLC
Figure 2. Avalanche®-Everyday Transfection Reagent (“Everyday”) and Avalanche®-Omni Transfection Reagent (“Omni”) achieved more than 90% knockdown of endogenous gene expression. (2A). ROCK1 mRNA levels were quantified using qPCR in Hela cells transfected with Target-specific siRNA duplexes (10nM) for human ROCK1 gene or non-silencing siRNA by using “Omni”, “Everyday”, and Lipofectamine® 2000 (“L2K”). Data were normalized against the 18S rRNA signal. Control Samples were either mock-transected or untreated. Values are normalized to untreated sample. Data are means®®SD (n=3). (B) Actin stress fibers were stained with FITC-labelled phalloidin on the cells in (A) cultured on fibronectin-coated micropattern. Confocal Microscope revealed that the cells transfected with ROCK1 siRNA showed disrupted stress fiber pattern as compared to the cells transfected with non-silencing siRNA using the same transfection reagent “Omni”. The use of “Everyday” showed similar disrupted stress fiber pattern as that of “Omni”, while the use of “L2K” only showed slightly disrupted stress fiber pattern. Lipofectamine® is a trademark of Life Technologies Corporation. Avalanche® is a trademark of EZ Biosystems® LLC
Additional Information
| Weight | 0.5 lbs |
|---|---|
| Product Sizes | 0.75 ml, 1.5 ml |
| Subcategories | Broad Spectrum |
Documents
Protocols
MSDS
Citations or Feedback
-
-
- Agbaegbu Iweka, C., Hussein, R. K., Yu, P., Katagiri, Y., & Geller, H. M. (2021). The lipid phosphatase-like protein PLPPR1 associates with RhoGDI1 to modulate RhoA activation in response to axon growth inhibitory molecules. Journal of Neurochemistry, 157(3), 494–507. https://doi.org/10.1111/jnc.15271
- Aso, S., Kitao, K., Hashimoto-Gotoh, A., Sakaguchi, S., & Miyazawa, T. (2021). Identification of feline foamy virus-derived microRNAs. Microbes and Environments, 36(4), Article ME21055. https://doi.org/10.1264/jsme2.ME21055
- Byrne, D. J., Garcia-Pardo, M. E., Cole, N. B., Stadler, J., Langton, G. M., Geller, H. M., & Blackstone, C. (2022). Liver X receptor-agonist treatment rescues degeneration in a Drosophila model of hereditary spastic paraplegia. Acta Neuropathologica Communications, 10, Article 40. https://doi.org/10.1186/s40478-022-01343-6
- Chai, P., Perr, J., Kageler, L., Lebedenko, C. G., Dias, J. M. L., Yankova, E., Esko, J. D., Tzelepis, K., & Flynn, R. A. (2024). Cell surface ribonucleoproteins cluster with heparan sulfate to regulate growth factor signaling [Preprint]. bioRxiv. https://doi.org/10.1101/2024.07.25.605163
- Chang, J., Lee, S., & Blackstone, C. (2014). Spastic paraplegia proteins spastizin and spatacsin mediate autophagic lysosome reformation. Journal of Clinical Investigation, 124(11), 5249–5262. https://doi.org/10.1172/JCI77598
- Chen, J., Harding, S. M., Natesan, R., Minn, A. J., Asangani, I. A., & Greenberg, R. A. (2020). Cell cycle checkpoints cooperate to suppress DNA- and RNA-associated molecular pattern recognition and anti-tumor immune responses. Cell Reports, 32(9), Article 108080. https://doi.org/10.1016/j.celrep.2020.108080
- Chesnel, F., Jullion, E., Delalande, O., Fautrel, A., Arlot-Bonnemains, Y., & Le Goff, X. (2022). Mutation of the proline P81 into a serine modifies the tumour suppressor function of the von Hippel–Lindau gene in the ccRCC. British Journal of Cancer, 127(11), 1954–1962. https://doi.org/10.1038/s41416-022-01985-2
- Chu, Y., Cohen, B. E., & Chuang, H.-h. (2020). A single TRPV1 amino acid controls species sensitivity to capsaicin. Scientific Reports, 10, Article 8038. https://doi.org/10.1038/s41598-020-64584-2
- Ebihara, T., Masuda, A., Takahashi, D., Mon, H., Hino, M., Fan, X., Lee, J. M., & Kusakabe, T. (2021). Production of scFv, Fab, and IgG of CR3022 antibodies against SARS-CoV-2 using silkworm-baculovirus expression system. Molecular Biotechnology, 63(12), 1223–1234. https://doi.org/10.1007/s12033-021-00373-0
- Filippov-Levy, N., Davidson, B., & Reich, R. (2020). The biological role of the long non-coding RNA LINK-A in ovarian carcinoma. Anticancer Research, 40(12), 6677–6684. https://doi.org/10.21873/anticanres.14691
- Filippov-Levy, N., Reich, R., & Davidson, B. (2020). The biological and clinical role of the long non-coding RNA LOC642852 in ovarian carcinoma. International Journal of Molecular Sciences, 21(15), Article 5237. https://doi.org/10.3390/ijms21155237
- Garaeva, F., Nakajima, R., Tamai, S., et al. (2025). Ataxin-2 as a candidate blood biomarker for estimating disease status in cases of suspected glioblastoma recurrence. Brain Tumor Pathology. Advance online publication. https://doi.org/10.1007/s10014-025-00517-z
- Garcia, B. C. B., Mukai, Y., Tomonaga, K., & Horie, M. (2023). The hidden diversity of ancient bornaviral sequences from X and P genes in vertebrate genomes. Virus Evolution, 9(1), Article vead038. https://doi.org/10.1093/ve/vead038
- Greenberg, E., Hochberg-Laufer, H., Blanga, S., Kinor, N., & Shav-Tal, Y. (2019). Cytoplasmic DNA can be detected by RNA fluorescence in situ hybridization. Nucleic Acids Research, 47(18), Article e109. https://doi.org/10.1093/nar/gkz645
- Hakata, H., Takai, Y., Lee, J. M., Kusakabe, T., Satone, H., Shimasaki, Y., & Oshima, Y. (2023). Tributyltin-binding protein type 1 (fish acid glycoprotein) is a potential gatekeeper of ethinylestradiol action in fish. Comparative Biochemistry and Physiology Part C: Toxicology & Pharmacology, 271, Article 109660. https://doi.org/10.1016/j.cbpc.2023.109660
- Hernandez-Vicens, R., Singh, J., Pernicone, N., Listovsky, T., & Gerlitz, G. (2022). SETDB1 regulates microtubule dynamics. Cell Proliferation, 55(12), Article e13348. https://doi.org/10.1111/cpr.13348
- Higashi, S. L., Yagyu, K., Nagase, H., Pearson, C. S., Geller, H. M., & Katagiri, Y. (2020). Old but not obsolete: An enhanced high-speed immunoblot. The Journal of Biochemistry, 168(1), 15–22. https://doi.org/10.1093/jb/mvaa016
- Hirakawa, T., Taniuchi, M., Iguchi, Y., Akashi, M., Imai, K., Matsuoka, S., Iida, S., Tomoike, K., Iizumi, Y., & Higuchi, T. (2025). NF-E2-related factor 1 suppresses the expression of a spermine oxidase and the production of highly reactive acrolein. Scientific Reports, 15, Article 12405. https://doi.org/10.1038/s41598-025-96388-7
- Iosefson, O., Olivares, A. O., Baker, T. A., & Sauer, R. T. (2015). Dissection of axial-pore loop function during unfolding and translocation by a AAA+ proteolytic machine. Cell Reports, 12(6), 1032–1041. https://doi.org/10.1016/j.celrep.2015.07.007
- Ito, T., Saito, A., Kamikawa, Y., Nakazawa, N., & Imaizumi, K. (2024). AIbZIP/CREB3L4 promotes cell proliferation via the SKP2-p27 axis in luminal androgen receptor subtype triple-negative breast cancer. Molecular Cancer Research, 22(4), 373–385. https://doi.org/10.1158/1541-7786.MCR-23-0629
- Iweka, C. A., Tilve, S., Mencio, C., Katagiri, Y., & Geller, H. M. (2019). The lipid phosphatase-like protein PLPPR1 increases cell adhesion by modulating RhoA/Rac1 activity [Preprint]. bioRxiv. https://doi.org/10.1101/470914
- Jiang, H., Zhang, T., Kaur, H., Kwon, Y., Sung, P., & Greenberg, R. A. (2024). BLM helicase unwinds lagging strand substrates to assemble the ALT telomere damage response. Molecular Cell, 84(9), 1684–1698.e9. https://doi.org/10.1016/j.molcel.2024.04.008
- Jo, T., Tsujimoto, K., Nakatani, T., Nagira, D., Muto, Y., Hirayama, T., Konaka, H., Okada, M., Takamatsu, H., & Kumanogoh, A. (2025). Lysosomal Ca²⁺ drives Lamtor1–MPRIP-mediated cell migration. Life Science Alliance, 8(8), Article e202403015. https://doi.org/10.26508/lsa.202403015
- Kakasaki, N., Koya, Y., Yamashita, M., Jo, T., Tsujimoto, K., Nakatani, T., Nagira, D., Muto, Y., Hirayama, T., & Konaka, H. (2024). GALNTL5, which is restricted to mouse spermatids, impairs endoplasmic reticulum (ER) function through direct interaction with ER chaperone proteins. Cell Death Discovery, 10, Article 499. https://doi.org/10.1038/s41420-024-02252-4
- Kamikawa, Y., Wu, Z., Nakazawa, N., Saito, A., & Imaizumi, K. (2023). Impact of cell cycle on repair of ruptured nuclear envelope and sensitivity to nuclear envelope stress in glioblastoma. Cell Death Discovery, 9, Article 233. https://doi.org/10.1038/s41420-023-01534-7
- Kang, J., Kim, J. W., Heo, H., Lee, J., Park, K. Y., Yoon, J. H., & Chang, J. (2021). Identification of BAG2 and Cathepsin D as plasma biomarkers for Parkinson’s disease. Clinical and Translational Science, 14(2), 606–616. https://doi.org/10.1111/cts.12920
- Kasahara, Y., Osuka, S., Takasaki, N., Jo, T., Tsujimoto, K., Nakatani, T., Nagira, D., Muto, Y., Hirayama, T., & Konaka, H. (2021). Primate-specific POTE-actin gene could play a role in human folliculogenesis by controlling the proliferation of granulosa cells. Cell Death Discovery, 7, Article 186. https://doi.org/10.1038/s41420-021-00566-1
- Katagiri, Y., Morgan, A. A., Yu, P., Bangayan, N. J., Junka, R., & Geller, H. M. (2018). Identification of novel binding sites for heparin in receptor protein-tyrosine phosphatase (RPTPσ): Implications for proteoglycan signaling. Journal of Biological Chemistry, 293(29), 11639–11647. https://doi.org/10.1074/jbc.RA118.004976
- Katagiri, Y., Morgan, A. A., Yu, P., Bangayan, N. J., Junka, R., & Geller, H. M. (2018). Identification of novel binding sites for heparin in receptor protein-tyrosine phosphatase (RPTPσ): Implications for proteoglycan signaling. Journal of Biological Chemistry, 293(29), 11639–11647. https://doi.org/10.1074/jbc.RA118.004976
- Kelly, C. M., Byrnes, L. J., Neela, N., Sondermann, H., & O’Donnell, J. P. (2021). The hypervariable region of atlastin-1 is a site for intrinsic and extrinsic regulation. Journal of Cell Biology, 220(11), Article e202104128. https://doi.org/10.1083/jcb.202104128
- Kim, J.-W., Jung, S.-Y., Kim, Y., Heo, H., Hong, C.-H., Seo, S.-W., Choi, S.-H., Son, S.-J., Lee, S., & Chang, J. (2021). Identification of Cathepsin D as a plasma biomarker for Alzheimer’s disease. Cells, 10(1), Article 138. https://doi.org/10.3390/cells10010138
- Kitao, K., Ichiyanagi, K., & Nakagawa, S. (2025). Birth of protein-coding exons by ancient domestication of LINE-1 retrotransposon. Genome Research, 35(12), 1287–1300. https://doi.org/10.1101/gr.280007.124
- Kitao, K., Nakagawa, S., & Miyazawa, T. (2021). An ancient retroviral RNA element hidden in mammalian genomes and its involvement in co-opted retroviral gene regulation. Retrovirology, 18, Article 36. https://doi.org/10.1186/s12977-021-00580-2
- Kitao, K., Shoji, H., Miyazawa, T., & Nakagawa, S. (2023). Dynamic evolution of retroviral envelope genes in egg-laying mammalian genomes. Molecular Biology and Evolution, 40(5), Article msad090. https://doi.org/10.1093/molbev/msad090
- Konaka, H., Kato, Y., Hirano, T., Jo, T., Tsujimoto, K., Nakatani, T., Nagira, D., Muto, Y., Hirayama, T., & Kumanogoh, A. (2023). Secretion of mitochondrial DNA via exosomes promotes inflammation in Behçet’s syndrome. The EMBO Journal, 42(13), Article e112573. https://doi.org/10.15252/embj.2022112573
- Kunitomi, A., Toyofuku, Y., Chiba, S., Higashitani, N., Higashitani, A., Sato, S., Mizuno, K., & Ohashi, K. (2025). Actin cytoskeletal remodeling requires the interaction between Solo and LARG in response to substrate stiffness [Preprint]. bioRxiv. https://doi.org/10.1101/2025.04.08.647765
- Lee, S., Chang, J., & Blackstone, C. (2016). FAM21 directs SNX27–retromer cargoes to the plasma membrane by preventing transport to the Golgi apparatus. Nature Communications, 7, Article 10939. https://doi.org/10.1038/ncomms10939
- Lee, S., Chang, J., & Blackstone, C. (2020). Hereditary spastic paraplegia SPG8 mutations impair CAV1-dependent, integrin-mediated cell adhesion. Science Signaling, 13(631), Article eaau7500. https://doi.org/10.1126/scisignal.aau7500
- Lenaers, G., Beignon, F., Ducreux, S., Tuifua, L., Le Dantec, Y., et al. (2024). A human mitochondrial isoform of TRPV1 regulates intracellular $Ca^{2+}$ simultaneously with mitochondrial thermolysis [Preprint]. Research Square. https://doi.org/10.21203/rs.3.rs-5111212/v1
- Lipinski, R. A. J., Stancill, J. S., Nuñez, R., Wynia-Smith, S. L., Sprague, D. J., Nord, J. A., Bird, A., Corbett, J. A., & Smith, B. C. (2024). Zinc-chelating BET bromodomain inhibitors equally target islet endocrine cell types. American Journal of Physiology-Regulatory, Integrative and Comparative Physiology, 326(6), R635–R647. https://doi.org/10.1152/ajpregu.00259.2023
- Liu, T.-Y., Chu, Y., Mei, H.-R., Chang, D. S., & Chuang, H.-H. (2019). It takes two vanilloid ligand bindings per channel to transduce painful capsaicin stimuli [Preprint]. bioRxiv. https://doi.org/10.1101/725598
- Liu, T.-Y., Chu, Y., Mei, H.-R., Chang, D. S., & Chuang, H.-H. (2020). Two vanilloid ligand bindings per channel are required to transduce capsaicin-activating stimuli. Frontiers in Molecular Neuroscience, 12, Article 302. https://doi.org/10.3389/fnmol.2019.00302
- Maeda, T. K., Sugiura, D., Okazaki, I.-m., Maruhashi, T., & Okazaki, T. (2019). Atypical motifs in the cytoplasmic region of the inhibitory immune co-receptor LAG-3 inhibit T cell activation. Journal of Biological Chemistry, 294(15), 6017–6026. https://doi.org/10.1074/jbc.RA119.007760
- Maruhashi, T., Okazaki, I.-m., Sugiura, D., Maeda, T. K., Shimizu, K., & Okazaki, T. (2018). LAG-3 inhibits the activation of CD4+ T cells that recognize stable pMHCII through its conformation-dependent recognition of pMHCII. Nature Immunology, 19(12), 1415–1426. https://doi.org/10.1038/s41590-018-0217-9
- Matsuyama, M., & Iwamiya, T. (2024). Novel and effective plasmid transfection protocols for functional analysis of genetic elements in human cardiac fibroblasts. PLOS ONE, 19(11), Article e0309566. https://doi.org/10.1371/journal.pone.0309566
- Morio, A., Lee, J. M., Fujii, T., Mon, H., Masuda, A., Kakino, K., Xu, J., Banno, Y., & Kusakabe, T. (2023). The biological role of core 1β1-3galactosyltransferase (T-synthase) in mucin-type O-glycosylation in silkworm, Bombyx mori. Insect Biochemistry and Molecular Biology, 156, Article 103936. https://doi.org/10.1016/j.ibmb.2023.103936
- Nakai, N., Sato, K., Tani, T., Kawagishi, M., Ka, H., Saito, K., & Terada, S. (2021). Development of nanobody-based POLArIS orientation probes enabled multi-color/multi-target orientation imaging in living cells. Biochemical and Biophysical Research Communications, 565, 50–56. https://doi.org/10.1016/j.bbrc.2021.05.088
- Nakai, N., Sato, K., Tani, T., Saito, K., Sato, F., & Terada, S. (2019). Genetically encoded orientation probes for F-actin for fluorescence polarization microscopy. Microscopy, 68(5), 359–368. https://doi.org/10.1093/jmicro/dfz022
- Nakamura, T., Hara, N., Osago, H., Hiyoshi, M., Kobayashi-Miura, M., & Tsuchiya, M. (2021). Effects of NAD+ synthesis levels on sirtuin 1 deacetylase activity in mammalian cells. Shimane Journal of Medical Science, 38(2), 59–66. https://doi.org/10.51010/sjms.38.2_59
- Nakatani, T., Tsujimoto, K., Park, J., Jo, T., Nagira, D., Muto, Y., Hirayama, T., Konaka, H., Okada, M., Takamatsu, H., & Kumanogoh, A. (2021). The lysosomal Ragulator complex plays an essential role in leukocyte trafficking by activating myosin II. Nature Communications, 12, Article 3333. https://doi.org/10.1038/s41467-021-23654-3
- Nasuhidehnavi, A., Zhao, Y., Punetha, A., Hemphill, A., Li, H., Bechtel, T. J., Rager, T., Xiong, B., Petrou, V. I., Gubbels, M.-J., Weerapana, E., & Yap, G. S. (2022). A role for Basigin in Toxoplasma gondii infection. Infection and Immunity, 90(6), Article e00205-22. https://doi.org/10.1128/iai.00205-22
- Nezich, C. L., Wang, C., Fogel, A. I., & Youle, R. J. (2015). MiT/TFE transcription factors are activated during mitophagy downstream of Parkin and Atg5. Journal of Cell Biology, 210(3), 435–450. https://doi.org/10.1083/jcb.201501002
- Ninomiya, H., Fukuda, S., Nishida-Fukuda, H., Shibata, Y., Sato, T., Nakamichi, Y., Nakamura, M., Udagawa, N., Miyazawa, K., & Suzuki, T. (2025). Osteoprotegerin secretion and its inhibition by RANKL in osteoblastic cells visualized using bioluminescence imaging. Bone, 191, Article 117319. https://doi.org/10.1016/j.bone.2024.117319
- Nohara, T., Kumamoto, J., Mai, Y., Shimano, M., Kato, S., Kitahata, H., Nakamura, H., Takashima, S., Watanabe, M., Nagayama, M., Oikawa, T., Ujiie, H., & Natsuga, K. (2024). Spatial confinement induces reciprocating migration of epidermal keratinocytes and forms triphasic epithelia [Preprint]. bioRxiv. https://doi.org/10.1101/2024.11.12.623158
- Nord, J. A., Wynia-Smith, S. L., Gehant, A. L., Jones Lipinski, R. A., Naatz, A., Rioja, I., Prinjha, R. K., Corbett, J. A., & Smith, B. C. (2022). N-terminal BET bromodomain inhibitors disrupt a BRD4-p65 interaction and reduce inducible nitric oxide synthase transcription in pancreatic β-cells. Frontiers in Endocrinology, 13, Article 923925. https://doi.org/10.3389/fendo.2022.923925
- Nuñez, R., Sidlowski, P. F. W., Steen, E. A., Wynia-Smith, S. L., Sprague, D. J., Keyes, R. F., & Smith, B. C. (2024). The TRIM33 bromodomain recognizes histone lysine lactylation. ACS Chemical Biology, 19(12), 2736–2743. https://doi.org/10.1021/acschembio.4c00789
- Oda, T., Yanagisawa, H., Arai, H., Miyazawa, H., et al. (2024). Cryo-EM of PMEL amyloids reveals pathogenic mechanism of pigment dispersion syndrome [Preprint]. Research Square. https://doi.org/10.21203/rs.3.rs-5540188/v1
- Ohga, H., Ito, K., Kakino, K., Mon, H., Kusakabe, T., Lee, J. M., & Matsuyama, M. (2021). Leptin is an important endocrine player that directly activates gonadotropic cells in teleost fish, chub mackerel. Cells, 10(12), Article 3505. https://doi.org/10.3390/cells10123505
- Onallah, H., Mannully, S. T., Davidson, B., & Reich, R. (2022). Exosome secretion and epithelial-mesenchymal transition in ovarian cancer are regulated by phospholipase D. International Journal of Molecular Sciences, 23(21), Article 13286. https://doi.org/10.3390/ijms232113286
- Osborn, J. L., & Greer, S. F. (2015). Metastatic melanoma cells evade immune detection by silencing STAT1. International Journal of Molecular Sciences, 16(2), 4343–4361. https://doi.org/10.3390/ijms16024343
- Park, H., Heo, H., Song, Y., Lee, M. S., Cho, Y., Lee, J.-S., Chang, J., & Lee, S. (2025). TRIM22 functions as a scaffold protein for autophagy initiation. Animal Cells and Systems, 29(1), 296–311. https://doi.org/10.1080/19768354.2025.2498926
- Renvoisé, B., Malone, B., Falgairolle, M., Munasinghe, J., Stadler, J., Sibilla, C., Park, S. H., & Blackstone, C. (2016). Reep1 null mice reveal a converging role for hereditary spastic paraplegia proteins in lipid droplet regulation. Human Molecular Genetics, 25(23), 5111–5125. https://doi.org/10.1093/hmg/ddw315
- Saito, A., Kamikawa, Y., Ito, T., Matsushita, Y., Katagiri, T., & Imaizumi, K. (2023). p53-independent tumor suppression by cell-cycle arrest via CREB/ATF transcription factor OASIS. Cell Reports, 42(5), Article 112479. https://doi.org/10.1016/j.celrep.2023.112479
- Satoh, S., Miyake, K., Adachi, Y., Masuhiro, K., Futami, S., Naito, Y., Shiroyama, T., Koyama, S., Yamaguchi, Y., Konaka, H., Takamatsu, H., Okuzaki, D., Nagatomo, I., Takeda, Y., & Kumanogoh, A. (2024). Cancer-associated SNRPD3 mutation confers resistance to hypoxia, which is attenuated by DRP1 inhibition. Biochemical and Biophysical Research Communications, 696, Article 149511. https://doi.org/10.1016/j.bbrc.2023.149511
- Sherry, J., Dolat, L., McMahon, E., Swaney, D. L., Bastidas, R. J., Johnson, J. R., Valdivia, R. H., Krogan, N. J., Elwell, C. A., & Engel, J. N. (2022). Chlamydia trachomatis effector Dre1 interacts with dynactin to reposition host organelles during infection [Preprint]. bioRxiv. https://doi.org/10.1101/2022.04.15.488217
- Shoji, H., Kitao, K., Miyazawa, T., & Nakagawa, S. (2023). Potentially reduced fusogenicity of syncytin-2 in New World monkeys. FEBS Open Bio, 13(1), Article 13555. https://doi.org/10.1002/2211-5463.13555
- Sugimoto, H., Fukuda, S., Yokawa, S., Hori, M., Ninomiya, H., Sato, T., Miyazawa, K., Kawai, T., Furuno, T., Inouye, S., Goto, S., & Suzuki, T. (2022). Visualization of osteocalcin and bone morphogenetic protein 2 (BMP2) secretion from osteoblastic cells by bioluminescence imaging. Biochemical and Biophysical Research Communications, 625, 123–130. https://doi.org/10.1016/j.bbrc.2022.10.042
- Sugita, Y., Hirai, Y., Goto, S. H., Tomonaga, K., Noda, T., & Horie, M. (2025). Structure and assembly of Borna disease virus 1 nucleoprotein–RNA complexes [Preprint]. bioRxiv. https://doi.org/10.1101/2025.05.26.654368
- Sugiura, D., Okazaki, I.-m., Maeda, T. K., Maruhashi, T., Shimizu, K., Yamamoto, R., Maki, A., Yasuda, H., Sugawara, S., & Okazaki, T. (2022). PD-1 agonism by anti-CD80 inhibits T cell activation and alleviates autoimmunity. Nature Immunology, 23(3), 399–410. https://doi.org/10.1038/s41590-021-01125-7
- Takatsuka, R., Ito, S., Iwai, S., & Kuraoka, I. (2017). An assay to detect DNA-damaging agents that induce nucleotide excision-repairable DNA lesions in living human cells. Mutation Research/Genetic Toxicology and Environmental Mutagenesis, 820, 1–7. https://doi.org/10.1016/j.mrgentox.2017.05.009
- Takanezawa, Y., Nakamura, R., Matsuda, H., Yagi, T., Egawa, Z., Sone, Y., Uraguchi, S., Adachi, T., & Kiyono, M. (2019). Intracellular demethylation of methylmercury to inorganic mercury by organomercurial lyase (MerB) strengthens cytotoxicity. Toxicological Sciences, 170(2), 438–451. https://doi.org/10.1093/toxsci/kfz094
- Tawarayama, H., Feng, Q., Murayama, N., Suzuki, N., & Nakazawa, T. (2019). Cyclin-dependent kinase inhibitor 2b mediates excitotoxicity-induced death of retinal ganglion cells. Investigative Ophthalmology & Visual Science, 60(13), 4479–4488. https://doi.org/10.1167/iovs.19-27396
- Tawarayama, H., Suzuki, N., Inoue-Yanagimachi, M., Himori, N., Tsuda, S., & Sato, K. (2020). Glutathione trisulfide prevents lipopolysaccharide-induced inflammatory gene expression in retinal pigment epithelial cells. Ocular Immunology and Inflammation, 28(5), 789–800. https://doi.org/10.1080/09273948.2020.1833224
- Terasawa, K., Kato, Y., Ikami, Y., Sakamoto, K., Ohtake, K., Kusano, S., Tomabechi, Y., Kukimoto-Niino, M., Shirouzu, M., Guan, J.-L., Kobayashi, T., Iwata, T., Watabe, T., Yokoyama, S., & Hara-Yokoyama, M. (2021). Direct homophilic interaction of LAMP2A with the two-domain architecture revealed by site-directed photo-crosslinks and steric hindrances in mammalian cells. Autophagy, 17(12), 4286–4304. https://doi.org/10.1080/15548627.2021.1911017
- Tilve, S., Iweka, C. A., Bao, J., Hawken, N., Mencio, C. P., & Geller, H. M. (2020). Phospholipid phosphatase related 1 (PLPPR1) increases cell adhesion through modulation of Rac1 activity. Experimental Cell Research, 389(2), Article 111911. https://doi.org/10.1016/j.yexcr.2020.111911
- Toyotome, T., Takahashi, H., & Kamei, K. (2016). MEIS3 is repressed in A549 lung epithelial cells by deoxynivalenol and the repression contributes to the deleterious effect. The Journal of Toxicological Sciences, 41(1), 25–31. https://doi.org/10.2131/jts.41.25
- Tsujimoto, K., Jo, T., Nagira, D., Nakatani, T., Muto, Y., Hirayama, T., Konaka, H., Okada, M., Takamatsu, H., & Kumanogoh, A. (2023). The lysosomal Ragulator complex activates NLRP3 inflammasome in vivo via HDAC6. The EMBO Journal, 42(14), Article e111389. https://doi.org/10.15252/embj.2022111389
- Tsujioka, S., Kodera, N., Maruyama, T., Noshiro, D., Toyoshima, Y. Y., & Ando, T. (2023). Imaging single CaMKII holoenzymes at work by high-speed atomic force microscopy. Science Advances, 9(36), Article eadh1069. https://doi.org/10.1126/sciadv.adh1069
- Wu, Z., Omura, I., Saito, A., Imaizumi, K., & Kamikawa, Y. (2024). VPS4B orchestrates response to nuclear envelope stress by regulating ESCRT-III dynamics in glioblastoma. Nucleus, 15(1), Article 2423660. https://doi.org/10.1080/19491034.2024.2423660
- Yamazaki, M., Maruyama, S., Abé, T., Tsuneki, M., Kato, H., Izumi, K., Tanuma, J.-i., Cheng, J., & Saku, T. (2020). Rac1-dependent phagocytosis of apoptotic cells by oral squamous cell carcinoma cells: A possible driving force for tumor progression. Experimental Cell Research, 392(1), Article 112013. https://doi.org/10.1016/j.yexcr.2020.112013
- Yanagisawa, H., Arai, H., Wang, T., Oda, T., Miyazawa, H., et al. (2025). Cryo-EM of wild-type and mutant PMEL amyloid cores reveals structural mechanism of pigment dispersion syndrome. Nature Communications, 16, Article 5411. https://doi.org/10.1038/s41467-025-61233-y
- Yonemoto, E., Ihara, R., Tanaka, E., & Mitani, T. (2024). Cocoa extract induces browning of white adipocytes and improves glucose intolerance in mice fed a high-fat diet. Bioscience, Biotechnology, and Biochemistry, 88(10), 1188–1198. https://doi.org/10.1093/bbb/zbae105
- Zheng, P., Obara, C. J., Szczesna, E., et al. (2022). Publisher Correction: ER proteins decipher the tubulin code to regulate organelle distribution. Nature, 604, Article E11. https://doi.org/10.1038/s41586-022-04656-7
-


